Research
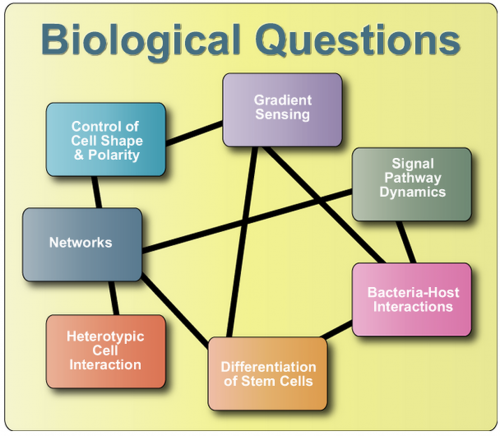
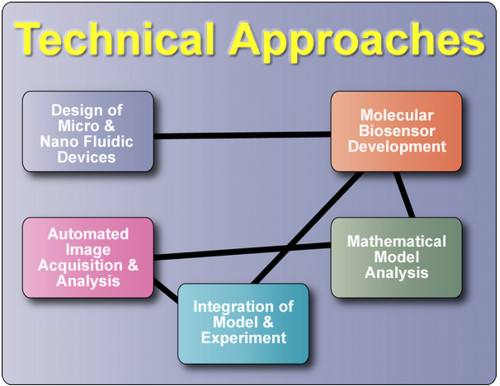
Sort research by Biological Questions - Sort research by Technical Approaches
Signaling Pathway Dynamics
Gradient Sensing
Traditionally, gradient sensing studies have been affected by lack of appropriate control of spatial (the actual shape of the gradient) or temporal gradients (gradient stability). Our lab has been developing a range of novel microfluidic device designs, which are tailor-made to allow accurately controlled gradient sensing studies for different cell-types. In addition, these studies are complemented by molecular biology approaches and mathematical modeling of the signal transduction pathways to understand the underlying basis of gradient sensing in these cells.
We study a diverse range of gradient sensing systems, ranging from pheromone sensing in yeast and axon guidance in embryonic Xenopus spinal neurons to the mammalian gradient sensing pathways involved in angiogenesis, tumor metastasis, haptotaxis and durotaxis.
Bacteria-Host Interactions
Differentiation of Stem Cells
Control of Cell Shape & Polarity
Heterotypic Cell Interaction
Cell behavior and functions are affected by its environment, which includes neighboring cells. This effect can be done by contact or by secreting soluble molecules. We investigate angiogenesis and tumor metastasis by studying interaction between tumor cells and endothelial cells for a wide range of scales: from single cell, to a group of cells to colonies. We are also interested in interactions between neuron and muscle cells, stem cell and differentiated cells, etc.
Networks
Biological networks are graphical representations of complex systems and processes, such as gene regulation, metabolism, or signal transduction. Large-scale topological analysis of these networks has revealed that they are not randomly organized. The intricate relations between their structure, function and dynamics need to be further investigated.
We study the dependence and interplay between network topology and the dynamics of the biological processes within a cell. We investigate ways in which network topology may have evolved to optimize the robustness of network dynamics. In addition, we investigate the notion of a close interplay between different kinds of biological networks: Specifically, we analyze how they interact as the cell receives and processes information about changes in its environment, and responds to them in optimal ways. To this end, we extensively study both the large-scale topology of biological networks and the function and dynamics of smaller topological motifs within them.
In our studies, we apply mathematical models, sophisticated simulation and visualization techniques, as well as our knowledge of the biochemical principles underlying network function. Our experimental facilities and resources permit us to change a cell’s environment in a controlled way, thus targeting different regions within the underlying biological network. By observing both single cells and in cell colonies, we can analyze diverse aspects of network dynamics.
Design of Micro & Nanofluidic Devices
Intracellular signal transduction involves a variety of molecular-level events occurring dynamically in space and time. To understand the signaling processes that, when integrated, generate specific cellular response, one often needs to perform multiple experimental perturbations of signaling network. The low throughput of traditional manual experimental technique is the limiting bottleneck in the collection of data necessary for constructing quantitative models of signaling transduction. Commercial “robotics” is too expensive and space consuming for typical laboratories, and often only capable of performing a single task.
Automated Image Acquisition & Analysis
Integration of Model & Experiment
Mathematical Model Analysis
We use a wide variety of different mathematical approaches to address many aspects of the biological questions we are currently investigating. Systems of ordinary differential equations (ODEs) are used to predict the behavior of complex intracellular signaling pathways and validated through both conventional and novel experimental approaches. For biological questions that are spatially dependent, such as chemotaxis and axonal guidance, we use both finite element and finite volume approaches to determine the significance of the polarization of signaling molecules. Depending on the question at hand we have also utilized a finer scale analysis involving Monte Carlo simulations tracking individual interacting molecules or agent based modeling aimed at describing multiple interacting cells within a population. In addition, we also use network analysis to model regulation of gene transcription and validate results using molecular biosensors.